
A/Professor Berin Broughton
The n=1 Project [MAY 2026]
When you study a cell, you usually have to destroy it first. Principal Research Fellow Berin Boughton has spent his career building the tools to see biology without losing what makes it real.
What a molecule does depends entirely on where it sits. Which cell it occupies. Which neighbours surround it. What stress the tissue is under at this particular moment in time.
For most of the history of biochemistry, we have studied molecules without their geography — grinding tissue, dissolving cells, extracting contents, and in doing so eliminating the very context that determines how biology actually works.
Principal Research Fellow Berin Boughton has been changing that. Based in the Department of Ecological, Plant and Animal Sciences at La Trobe University, he is one of the key figures responsible for introducing and advancing spatial metabolomics in Australia — mapping exactly which molecules are present in a tissue, where they are, and how they are changing, directly within intact tissue itself.
His work spans barley roots under cold stress and mouse eyes under metabolic strain. Quinoa crops and Huntington's disease. Endometriosis and parasitic drug resistance. What connects them is not a shared disease or a shared crop — it is a shared question, and a shared set of tools uniquely built to answer it.
This is a career that moved from organic chemistry to metabolomics to spatial 'omics — partly by design, partly by opportunity and luck, and always in the direction of problems that genuinely matter.
This is n=1.
We have been studying cells by destroying them.
THE PROBLEM
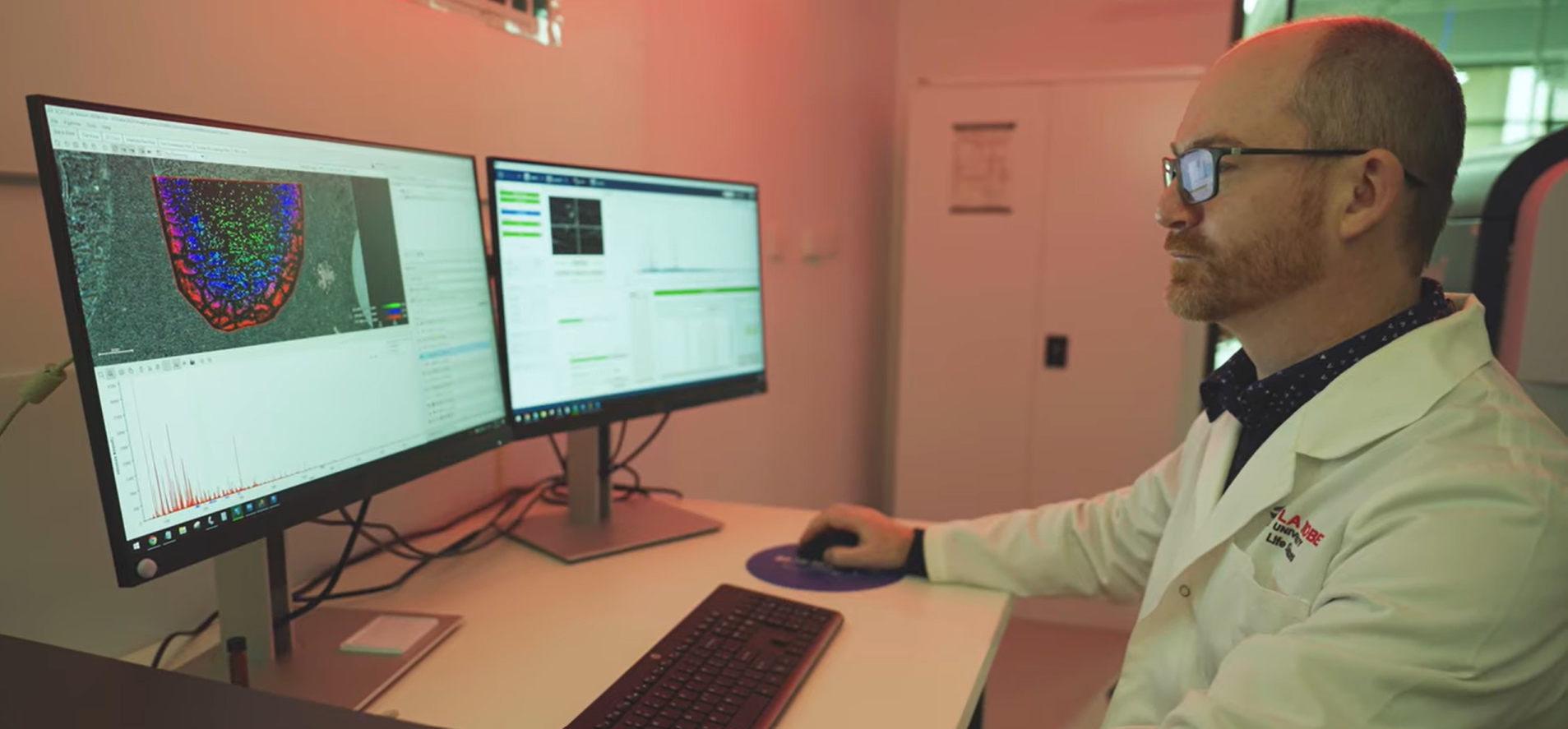

For decades, the standard approach to understanding molecular biology has been to break things apart. Grind tissue into homogenate. Extract molecules into solution. Measure what is present across the whole sample. This approach has been extraordinarily productive and underpins much of what we know about metabolism, disease, and drug response.
But it has a fundamental limitation.
Cells in complex organisms do not exist in isolation. They exist surrounded by other cell types, each performing distinct functions, each communicating constantly with their neighbours. That spatial context is not incidental to what a cell does. Which cell sits next to which, in which microenvironment, under what conditions: these are not background details. They are central to the biology itself.
Single-cell 'omics approaches have brought extraordinary resolution to molecular biology, but they strip away this spatial context entirely. Many imaging approaches preserve spatial information but lose the temporal dimension, the way molecular landscapes develop and shift over time. The result is a field that has long been forced to choose between resolution and context.
Spatial metabolomics and mass spectrometry imaging are beginning to make that trade-off unnecessary.
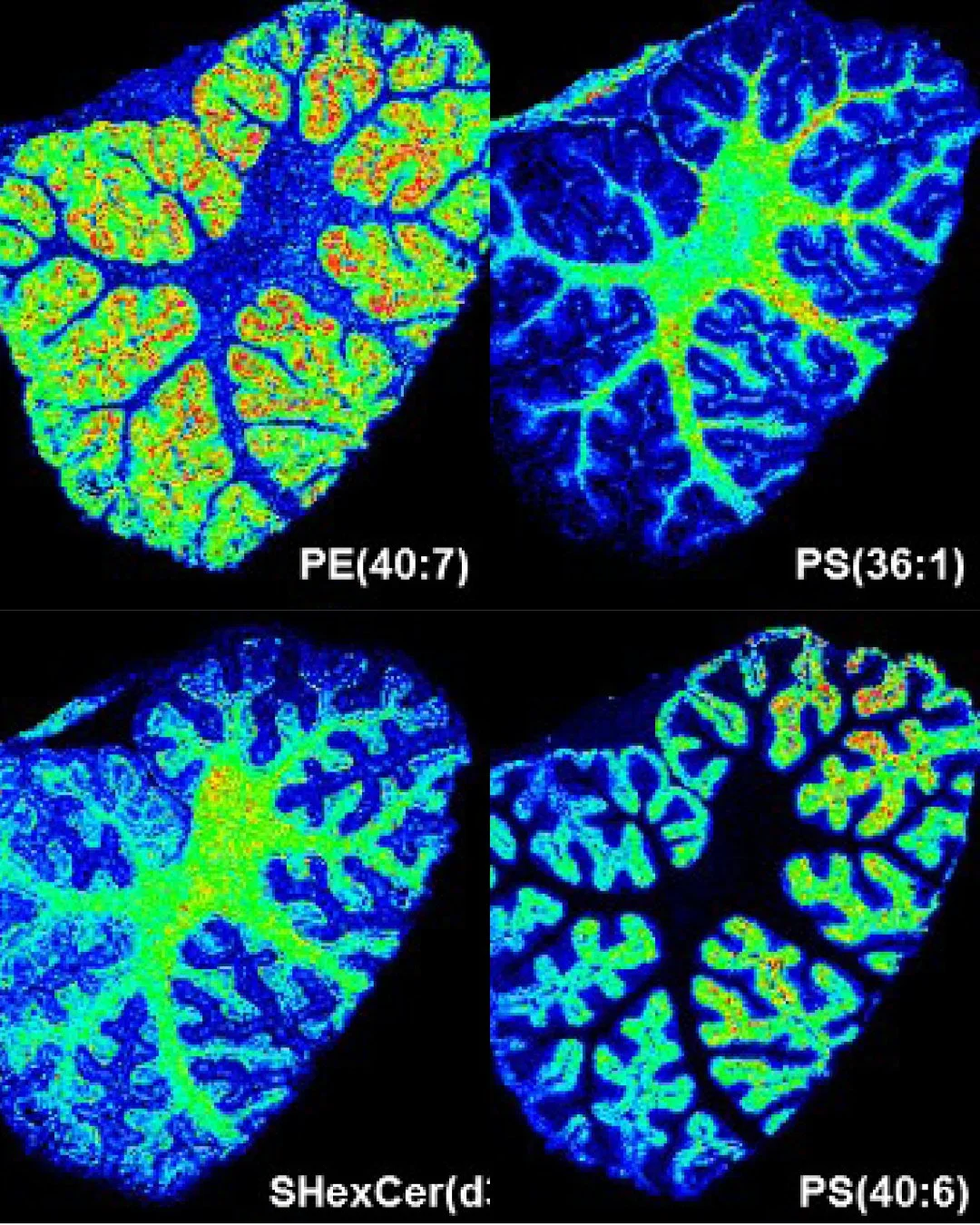
Seeing what was always there, but invisible.
Mass Spectrometry Imaging is sometimes described as molecular microscopy. Where a conventional microscope shows structure, it shows chemistry, mapping molecules directly from intact tissue with no labels required and no averaging across cell populations.
Berin's research group at La Trobe's Institute for Sustainable Agriculture and Food applies these tools across a remarkable breadth of biological problems.
THE RESEARCH
Plants, crops and food security
In plants, his team is examining what happens in barley root tissue when cold stress sets in: how the translational machinery responds, how the proteome is reshaped, and what this reveals about crop resilience mechanisms with direct relevance to Australian agriculture. This extends to quinoa, where Berin contributed to the first-ever mass spectrometry imaging of the grain, published in Nature in collaboration with researchers at KAUST, providing foundational resources for the genetic improvement of one of the world's most nutritious and food-secure crops.
Eyes, the brain and neurological disease
His team is mapping how distinct retinal cell types respond differently to the stress that drives diabetic retinopathy and macular degeneration. Earlier work using stable isotope labelling revealed pre-symptomatic changes in lipid metabolism in a Huntington's disease mouse model, identifying shifts in the hippocampus that preceded the onset of disease.
Endometriosis, atherosclerosis and infectious disease
Berin collaborates on endometriosis and atherosclerosis research, and his infectious disease work revealed that parasitic lesions contain metabolically quiescent parasites that survive drug treatment by switching to a near-dormant state, with broad implications for understanding drug resistance.
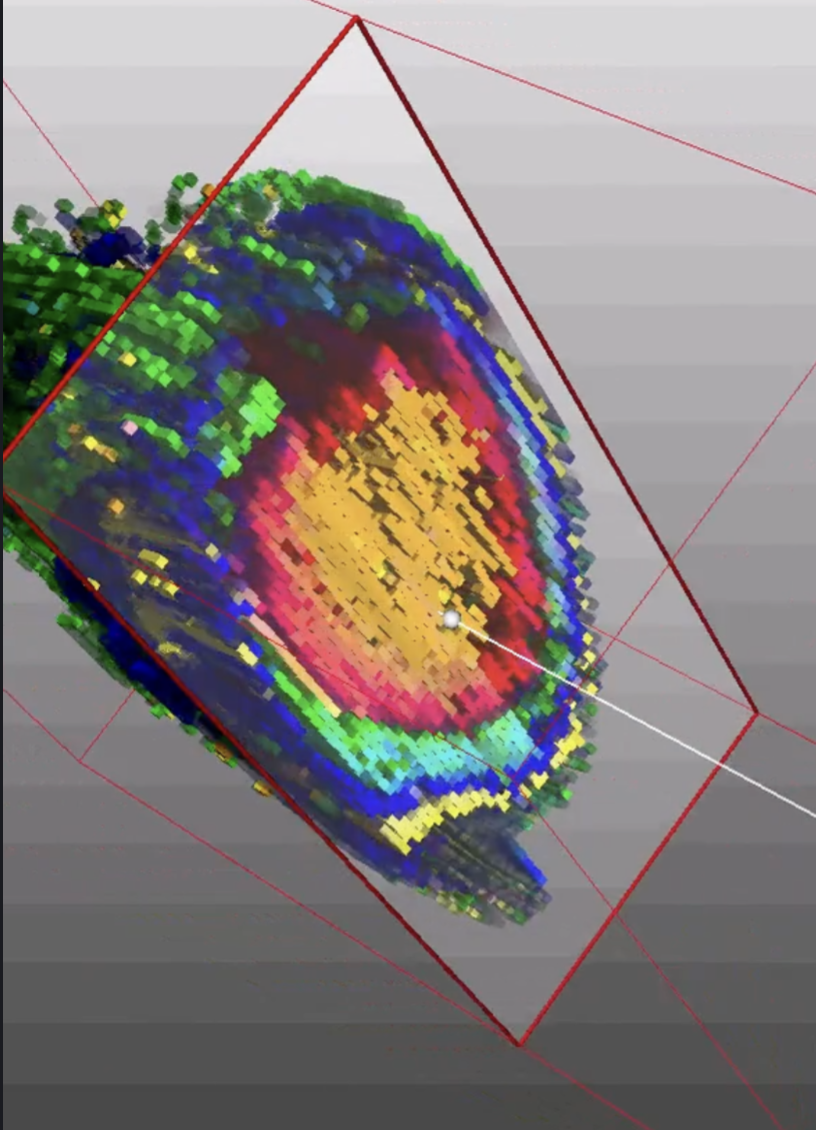


From organic chemistry to molecular geography.
Berin completed his PhD in chemistry at the University of Melbourne in 2010, with a background spanning organic chemistry and pharmacology. His early research included work on compounds designed to improve radiotherapy outcomes and reduce its severe side effects — a formative experience in applying technical chemistry to problems with direct human consequence.
THE JOURNEY
Joining Metabolomics Australia shortly before completing his PhD, Berin moved from service delivery into something closer to field-building. Over a decade, he introduced disciplines that had no established presence in Australian bioscience and built the infrastructure to sustain them.
His contributions across this period include:
Magnetic Resonance Mass Spectrometry — introduced and implemented in Australia for the first time
Spatial Metabolomics and Mass Spectrometry Imaging — pioneered nationally, with workflows, training and infrastructure built from the ground up
Amino acid analysis pipelines — generating more than a million dollars in revenue between 2011 and 2019
National team leadership — Deputy Head, then Head of Analytics, leading a team of more than ten researchers by 2018
Australian National Phenome Centre — ultra-high resolution magnetic resonance mass spectrometry focus from 2020 to 2023
METASPACE — contributor to the global metabolite annotation engine that has significantly accelerated MSI-based identification
MIAMSIE reporting standards — contributor to the international framework addressing reproducibility in mass spectrometry imaging research
In 2023 he joined La Trobe University to establish his own research group, bringing with him an established MSI facility and a collaborative network spanning Copenhagen, Auckland, Göttingen, the Max Planck Institute, and Vanderbilt University.
The motivation driving this trajectory has not changed, even as the tools and the targets have.
“What has driven me across more than two decades of research is a consistent desire to reduce suffering in the world — whether that's improving outcomes for cancer patients, developing new antibiotics, understanding diseases like endometriosis and atherosclerosis, or now addressing food security. The specific problems have changed, and the tools and expertise I've developed — partly by design, partly by opportunity and luck — have evolved considerably with them. But the underlying motivation hasn't. I'm drawn to technical science precisely because it gives you leverage — the ability to move the needle, even incrementally, on problems that genuinely matter.”
The missing dimension in biology
THE INSIGHT
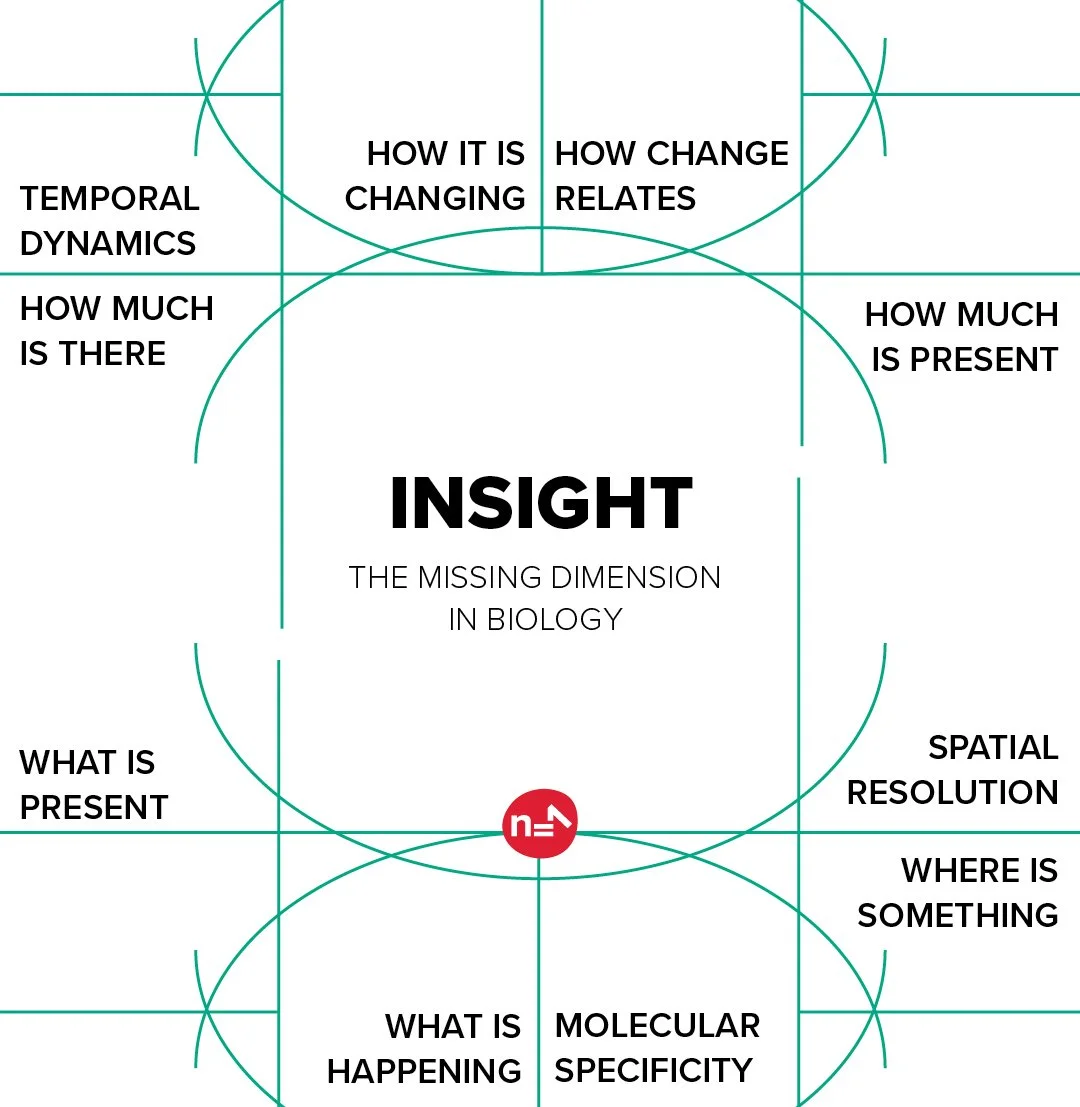
Most analytical approaches in biology resolve what is present, or how much. Far fewer tell you where. And very few are able to tell you where something is, how much of it there is, how it is changing, and how that change relates to what is happening in the cells immediately beside it.
That combination — spatial resolution, temporal dynamics, molecular specificity — is what spatial metabolomics, at its most developed, can provide. And it is what makes Berin's observation about conventional biology so pointed.
“Cells in complex organisms don't exist in isolation. They exist surrounded by different cell types, each performing distinct functions and communicating constantly to moderate and enhance each other's behaviour. That spatial and temporal context is fundamental to what each cell actually does — yet most of our current analytical approaches either destroy it or ignore it. Single-cell 'omics approaches are powerful, but they strip away context. Many imaging approaches preserve spatial information but lose the temporal dimension — how things develop and change. Our work is about recovering both simultaneously.”
THE IMPACT
What is visible now that was not visible before
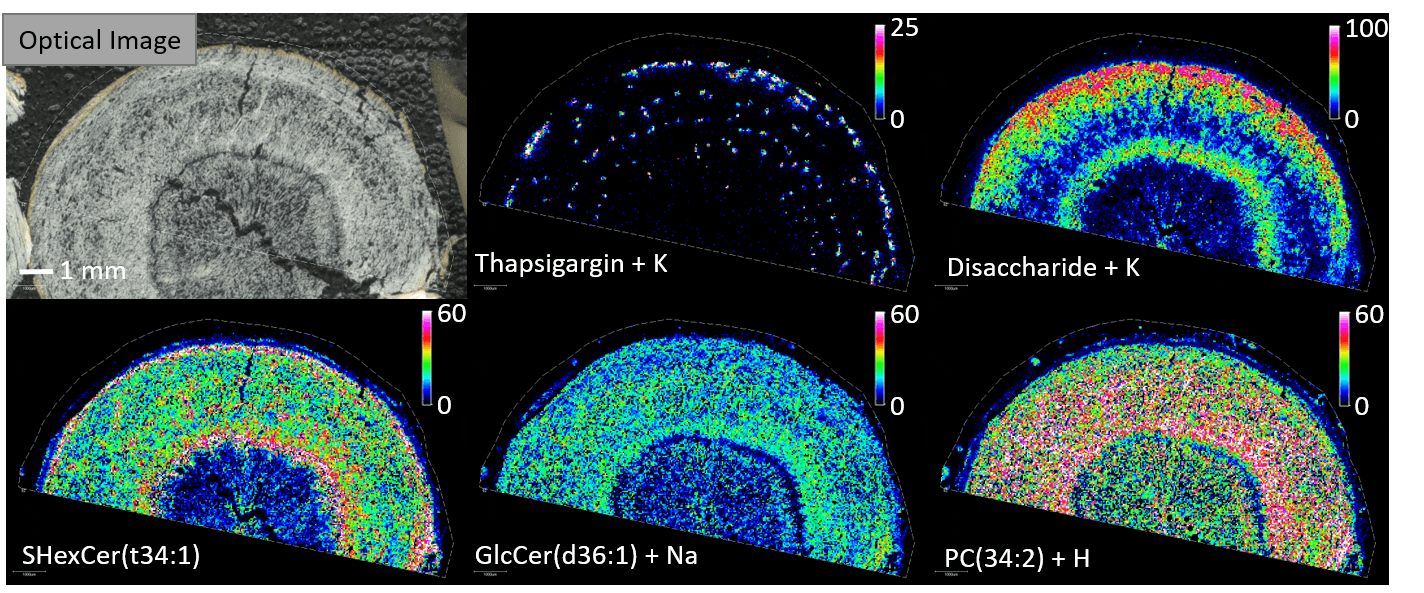
The breadth of Berin's research reflects something important about the nature of platform technologies. MSI and spatial metabolomics are not answers to specific biological questions — they are ways of asking questions more precisely. And that precision, applied across enough domains, becomes something larger than the sum of its parts.
In agriculture, the ability to map metabolic responses to cold stress in barley and trace molecular signatures in nitrogen-fixing root nodules offers a path toward crop varieties with enhanced resilience and reduced fertiliser dependence — outcomes with direct relevance to Australian food security at a moment when geopolitical disruption is already affecting global fertiliser supply.
In medicine, the capacity to spatially map lipid and metabolite dynamics in disease models — pre-symptomatically in Huntington's disease, at cellular resolution in the retina, at tissue level in endometriotic lesions — informs a new generation of therapeutic targets where current approaches are limited by the very methods used to generate the evidence.
In environmental science, minimally destructive spatial molecular mapping opens new possibilities for analysing rare, irreplaceable, or heritage specimens where conventional extraction would compromise the sample.
And in the development of Australian scientific infrastructure, Berin's current collaboration with Curtin University to build a next-generation MRT-enabled spatial chemistry platform represents a national investment in a capability that does not yet exist at scale anywhere in Australia.

“Fundamental understanding is often a prerequisite for applied solutions. Food security is no longer an abstract future concern — geopolitical disruption is already affecting fertiliser production and climate change is making frost events less predictable. I want the research my teams and collaborators are doing to have contributed molecular tools that plant breeders can actually use. But the same principles apply across systems — if the tools we're building together can illuminate stress responses across kingdoms, that's a contribution I'd be genuinely proud of.”

Dr Berin Broughton
n=-1 FEATURED RESEARCHER [MAY.26]
Read more about his research mapping the molecular landscape of living tissue across disease, agriculture, and beyond
Learn more about Berin's career and his role in establishing spatial metabolomics in Australia
To support the infrastructure and discovery research driving Berin's work, you can donate to La Trobe University here
Why n=1?
An n=1 study focuses deeply on one individual over time. Case studies provide practical, real-life insights that population studies cannot.
The same applies to research careers.
By presenting detailed observations of individual researchers, we demonstrate what excellence, dedication, and vision look like in practice. Not as abstract concepts or averaged data points, but as lived experience.
The intended outcomes:
For researchers: Recognition that your path doesn't need to mirror anyone else's to be valid. Your unique combination of talents and choices is the point.
For funders and policymakers: Evidence of the exceptional talent conducting groundbreaking research across Australia. Real people. Real projects. Real impact that deserves sustainable investment.
For the broader public: Understanding that research careers are as diverse as the problems they address. There is no template. There is only the unrepeatable intersection of expertise, passion, and opportunity that each researcher represents.
Your career is n=1.
Whether you've found your path inside or outside academia, whether your path looks like anyone else's or not, to validate your career you only need a sample size of one.
Yours.
